Student Project: Build an AI Plant Doctor App with Streamlit, PyTorch, and MongoDB Atlas
Moises Limon
What if diagnosing plant diseases were as easy as taking a photo? That's the idea behind 🌱 Dr. Plant, an AI-powered web app built to help farmers and gardeners identify plant illnesses and get treatment recommendations in seconds. Growing up with various tomato plants, lemon, and avocado trees getting sick, I have a personal connection to the struggles farmers face. Understanding the impact of plant diseases firsthand inspired me to develop a tool that could assist both small-scale gardeners and large agricultural producers. In this article, I'll walk through how I built Dr. Plant and how you can use a convolutional neural network (CNN), Google Gemini, Streamlit, PyTorch, and MongoDB Atlas for fast, scalable data storage.
What you'll build
You will build a web-based AI assistant that lets users upload images of plant leaves and returns:
The predicted plant disease
A tailored treatment plan (via Gemini)
Cached responses from MongoDB Atlas to boost performance

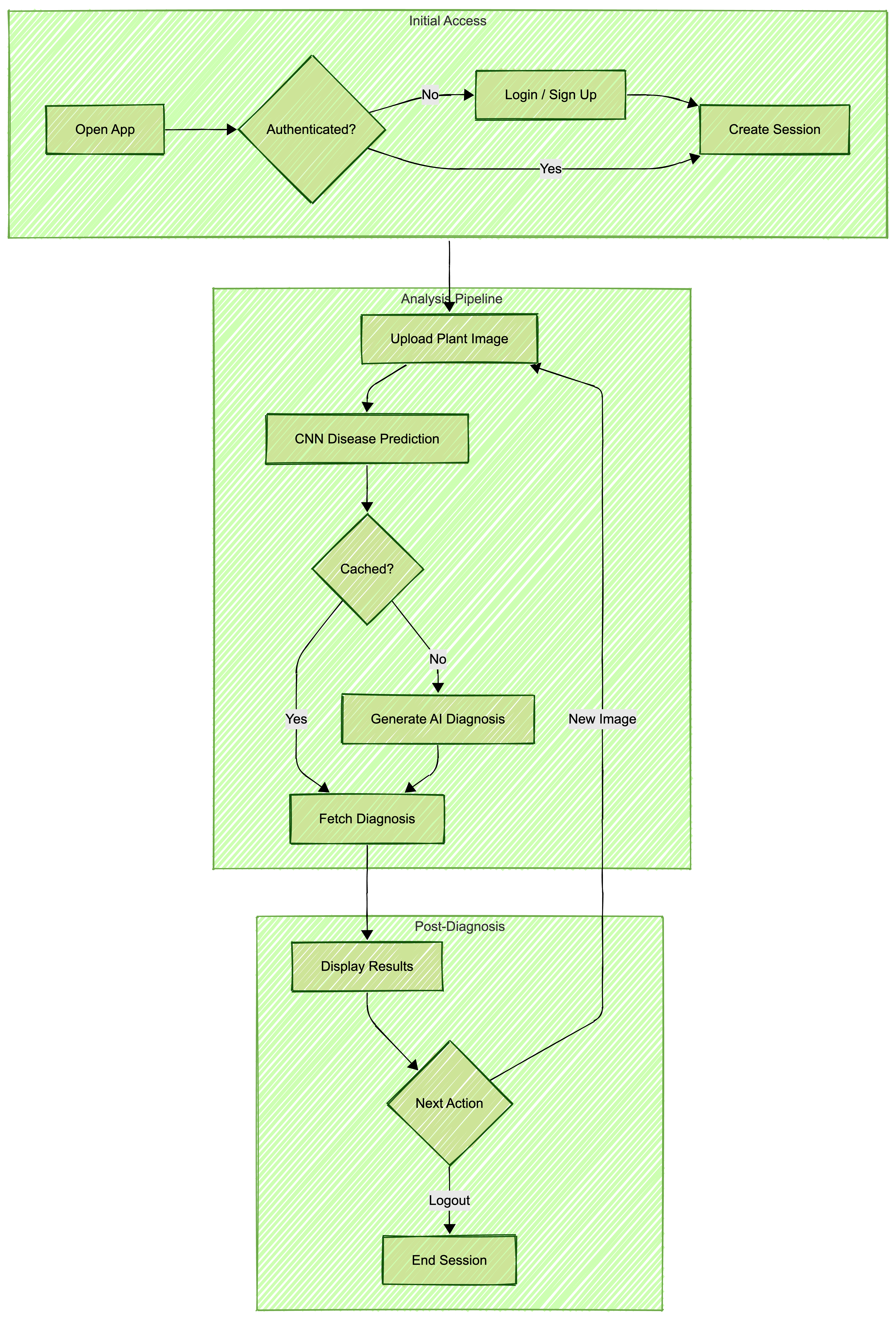
Application Architecture
The essential tools required to replicate this project include:
Python for model training and backend logic
PyTorch to build the CNN
Streamlit for the web UI
MongoDB Atlas to store image predictions and associated treatments
Google Gemini for treatment recommendations
PyMongo to connect Python to MongoDB
Dataset and model training
I used a publicly available dataset of plant leaf images labeled by disease. First I preprocessed the images by resizing them to 128x128 pixels, converting to normalized tensors, and adding random horizontal flips to create more data variation to help the model train better. I then trained a CNN model in Google Colab. For detailed steps on how to train this model, refer to the dev.ipynb in my GitHub repository.
Model architecture choices for the CNN
In order for the Dr. Plant app to efficiently classify plant diseases, the CNN was designed as follows:
Convolutional layers: Three layers with increasing filters (32, 64, 128) to capture complex patterns
Max pooling: Reduces feature map size, lowering computational load and overfitting risk
Fully connected layers: Two layers, first with 256 neurons for feature representation, second for class probabilities
Dropout: 0.5 rate to prevent overfitting by randomly deactivating neurons
ReLU activation: Introduces non-linearity, enabling complex pattern learning
These choices ensure accurate and efficient disease classification.
Model training and results
The CNN model was trained using the PyTorch framework, leveraging a dataset of plant leaf images. Here are the key details of the training process:
Dataset: The training set consisted of 70,295 images, while the validation set had 17,572 images, covering 38 different classes of plant diseases.
Training process: The model was trained over 10 epochs using the Adam optimizer with a learning rate of 0.001. The loss function used was CrossEntropyLoss since that is the standard loss function for multiclass classification tasks.
Epoch losses: The training loss decreased steadily over the epochs, starting from 1.2015 in the first epoch to 0.1565 by the tenth epoch.
Validation accuracy: The model achieved a validation accuracy of 96.82%, indicating its effectiveness in classifying plant diseases.
These results demonstrate the model's capability to accurately identify a wide range of plant diseases from leaf images. The model outputs a class label that identifies the plant disease.
Building the Streamlit app
Using Streamlit, I quickly created a frontend for users to upload images and receive diagnostic results. The complete implementation can be found here, but here's a simplified version of the prediction pipeline:
import streamlit as st
from PIL import Image
...
# Streamlit UI
st.title("🌱 Dr. Plant: Plant Disease Diagnosis AI")
uploaded_file = st.file_uploader("Upload an image of the plant", type=["jpg", "png", "jpeg"])
if uploaded_file:
image = Image.open(uploaded_file)
st.image(image, caption="Uploaded Image", width=600)Integrating MongoDB Atlas
To avoid redundant API calls and to speed up frequent queries, I used MongoDB Atlas to cache results. This approach not only reduces the load on the model but also provides a cost-effective solution by minimizing the number of model invocations. If a user submits an image previously seen, the app returns the result from MongoDB instantly, ensuring a fast response time and an enhanced user experience. The complete implementation can be found here, but here is the core logic summarized:
# Connect to MongoDB
client = pymongo.MongoClient("your_mongo_uri")
db = client["plant_doctor"]
disease_collection = db["plant_diseases"]
...
# Check if disease is already in MongoDB
existing_disease = disease_collection.find_one({"disease_label": disease_label})
if existing_disease:
diagnosis = existing_disease.get("disease_detail", "No diagnosis found.")
treatment = existing_disease.get("treatment_detail", "No treatment found.")
st.subheader("🩺 Diagnosis")
st.write(diagnosis.strip())
st.subheader("💊 Recommended Treatment")
st.write(treatment.strip())Getting treatment recommendations with Gemini
After identifying the disease, Dr. Plant queries Gemini to return a concise, user-friendly description and treatment plan (full implementation here). This integration allows responses to be tailored to each diagnosis in natural language.
# Query Gemini if disease is not found
model_gemini = genai.GenerativeModel(
"gemini-2.5-flash",
system_instruction=f"""
You are a plant disease expert.
Return ONLY valid JSON with exactly two fields:
- diagnosis
- treatment
Each field should be a detailed paragraph.
Explain why the disease is harmful in the diagnosis.
Treatment should include actionable steps and resources.
Do not include markdown or extra text
"""
)
try:
response = model_gemini.generate_content(
disease_label,
generation_config=genai.types.GenerationConfig(
temperature=0.7,
response_mime_type="application/json",
max_output_tokens=2000
)
)
...Implementing secure user login
Dr. Plant features a secure user login system to protect user data. Credentials are stored in MongoDB, with passwords encrypted using bcrypt, providing a cost-effective security solution. This method leverages open-source libraries and MongoDB's capabilities, avoiding the need for third-party services. Regular updates to encryption libraries are essential to maintain security. Future plans include integrating single sign-on (SSO) for enhanced user convenience, allowing access with existing credentials from other platforms. These measures ensure user data is handled with care, offering a safe and reliable experience.
Challenges and learnings
As with any development process, building Dr. Plant had its challenges and discoveries. Here is what I encountered and took away:
Streamlit components proved tricky when handling saved states and preserving session data across different user interactions.
Fine-tuning the CNN model to generalize well across multiple plant species required extensive experimentation.
Managing and structuring Gemini AI responses to ensure high-quality, actionable insights provided a valuable learning opportunity.
Future Enhancements
Looking ahead, here are some exciting enhancements you can make using Dr. Plant:
Deployment: Beyond the current UI implementation, refining the Streamlit application to utilize additional caching layers, whether that is through Streamlit’s native caching decorators or integrating Redis for distributed caching, would further improve response times and reduce database queries. Moreover, an additional caching layer would enable adding features such as a diagnosis history page, for users to quickly review their past diagnoses without repeated database lookups while reducing load on MongoDB .
Dataset expansion: Currently, the model is being trained on a fixed dataset. However, every user interaction with the application generates valuable data that could improve the model over time. Adding an opt-in feature for collecting user-submitted images alongside feedback on diagnosis accuracy can help the model to learn from a large variety of diseases. Open-sourcing this dataset would accelerate agricultural AI research, and the collective knowledge of plant disease detection would grow.
GCP integration: This prototype can be extended to be fully deployed in the Google Cloud environment. Using different GCP resources, this data can be stored in the GCP bucket for further analysis and scalability
Now you have all the information and knowledge you need to replicate this or build upon it! If you want to contribute or customize it, fork the repo, connect your own MongoDB Atlas cluster, and deploy on Streamlit Cloud or your platform of choice.
Let's plant a seed for innovation and cultivate AI systems that grow with efficiency!
Acknowledgements
I extend my gratitude to the USFCA MSDS program for all the learnings that cultivated this project coming to fruition, especially Dr. Mahesh Chaundhari for the opportunity. The Kaggle dataset used in this project is based on the work by Mohanty, Sharada P., Hughes, David P., and Salathé, Marcel, titled "Using deep learning for image-based plant disease detection," published in Frontiers in Plant Science, 2016. DOI: 10.3389/fpls.2016.01419.
Honoring Dr. Mahesh Chaudhari's Mentorship
Dr. Mahesh Chaudhari is an Assistant Professor in the Master of Science in Data Science and Artificial Intelligence (MS-DSAI) program and the Director of the Data Engineering program at the University of San Francisco. He is a driving force in data systems education. With 11 years of industry experience building scalable data infrastructure in multi-cloud environments as well as expertise in distributed databases, Dr. Chaudhari has a deep commitment to integrating MongoDB into data science and engineering curricula. His mentorship empowers students to create high-quality technical contributions, including innovative projects like Dr. Plant. His guidance bridges academic theory with real-world impact, elevating both MongoDB and student success.
Contact
For more questions or to discuss this project further, feel free to reach out to me via GitHub or LinkedIn. I welcome any feedback or collaboration opportunities.

